
The impact of translational studies in the current management of osteoporosis
Introduction
Recent technological and scientific advances have changed the way through which knowledge is produced. Until the early 1980s, research that was not hypothesis-oriented was considered an “anathema” within the scientific world. Yet, since the early 1990s, with the advent of high throughput methodologies, a new approach to scientific research was created, namely “discovery-driven” research. From these two opposite concepts another discipline arose, named “translational research” (TR). This term is often used interchangeably with translational medicine or translational science or bench to bedside and it is considered to be an effort to build on basic scientific research to create new therapies, medical procedures, or diagnostics (1). TR is essentially derived from the application of basic knowledge gained through basic research, which mainly identifies the mechanism of a disease or biological phenomenon. Basic knowledge gained in the laboratory when “translated” to potential treatments for a disease forms TR.
Osteoporosis is a systemic skeletal disorder characterized by decreased bone mass and a deterioration of bone microarchitecture, leading to enhanced fragility and a subsequent increase in fracture risk (2). It is also called “the silent disease”, as its presence often goes undetected until a low-energy fragility fracture occurs, after which a mechanism of diagnosis and management is put into place.
As a golden rule, bone strength has been clinically estimated in the recent decades non-invasively with the measurement of bone mineral density (BMD) by dual-energy X-ray absorptiometry (DEXA). The operational definition of osteoporosis proposed by the World Health Organization (WHO) is a BMD that lies 2.5 standard deviations (SD) or more below the average value for young healthy women (a T-score of <−2.5 SD) in post-menopausal women and men aged ≥50 years. Normal BMD is defined as a T score of −1.0 or higher; a T score between −1.0 and −2.5 is defined as osteopenia and a BMD T-score of ≤−2.5 SD associated with one or more fragility fractures defines established or severe osteoporosis (3).
Although this categorization identifies individuals with low BMD at increased risk of fracture, the majority of fragility fractures have been found to occur in individuals with osteopenia or even normal BMD, allegedly at lower risk with this method of estimation (4). Therefore, the need for the inclusion of additional methods, such as measurement of bone microarchitectural parameters with peripheral quantitative computerized tomography, or the inclusion of clinical risk factors in screening for the prevention of osteoporosis-related fractures, were identified. The latter led to the development of a number of tools, one of which is the WHO Fracture Risk Assessment Tool FRAX (5). Recent studies demonstrated that using the risk assessment tool in addition to BMD measurement in women aged 70–85 succeeds in identifying individuals at risk, reduces the incidence of hip fractures and is highly cost-effective.
Following diagnosis, management of osteoporosis includes pharmaceutical therapy and life-style modifications. Some of the antiresorptive therapeutic options are associated with side effects, decreasing patients’ compliance to drug therapy. As examples, estrogen replacement therapy is associated with increased cardiovascular incidents, pulmonary embolism and breast cancer; bisphosphonates are connected to gastrointestinal side effects, atypical femoral fractures and osteonecrosis of the jaw, even though the latter has a low incidence and depends on the concurrence of several factors. Therefore, it is frequently observed that non-compliant patients with osteoporosis often turn to alternative methods of therapy.
Hence, the present work attempts to outline some of the important concepts of TR in the treatment and the future of osteoporosis. Towards that purpose, we have reviewed the present knowledge on genomics and the use of in vitro and in vivo systems that are implemented towards the understanding of osteoporosis pathogenesis and treatment.
Translational studies in laboratory animals
Patients with low bone density unwilling to comply with conventional drug options show an increased interest in dietary consumption of plants, herbs or their extracts, which have demonstrated a bone-protective effect in translational studies. A number of these have been named “functional foods” and “medicinal plants”. Among their mechanisms of action are enhancement of osteoblast proliferation, antioxidant properties, mild phytoestrogenic action or modulation of the immune system (6), demonstrated mainly in the widely accepted ovariectomized rat model of osteoporosis (7). In this animal model, as estrogen withdrawal through ovariectomy results in statistically significant bone loss, which at the proximal tibial metaphysis is evident as early as 14 days postoperatively (8), many plant-derived substances have been administered in vivo to test their potentially beneficial action, either preventive or therapeutic.
Traditional Chinese medicinal plants have been used for generations in the prevention and treatment of osteoporosis. Current evidence-based experimental studies are confirming their bone beneficial effects, determining their mechanism of action and assessing their safety. The per os administration of naringin, a flavanone glucoside found in citrus fruits and in particular grapefruit, has been experimentally documented to be effective in a variety of disease models, such as atherosclerosis, cardiovascular disease, cancer models, diabetes, as well as osteoporosis (9). In the rat osteoporosis model, naringin increased BMD, bone volume/tissue volume, trabecular thickness and number, as well as femoral microvessel number in comparison to the untreated ovariectomized rats (10). Similarly, icariin administration has been documented to increase BMD, bone trabecular area and thickness, decrease trabecular separation and bone turnover markers, and increase bone biomechanical strength in comparison to the untreated ovariectomized rats (11). These effects result from icariin’s triggering the differentiation and mineralization of osteoblasts, while inhibiting osteoclast differentiation and inducing their apoptosis (12). Additionally, icariin treatment of ovariectomized rats resulted in significantly increased levels of β-catenin, Runx2 and Lrp6 mRNA compared to the untreated group, supporting that its bone-protective effect is mediated through the Wnt/β-catenin pathway (11).
Natural products such as green or black tea extracts administered to ovariectomized rats have also shown a beneficial effect on bone mass, cortical bone thickness, biomechanical properties and biomarkers of bone resorption (13). Other herb extracts from the Mediterranean region, such as Sideritis euboea and Onobrychis ebenoides, have also demonstrated beneficial effects on the estrogen-deficient rat skeleton by improving bone density, biomechanical properties and bone histomorphometry (6). Fresh juice from the kiwi fruit (Actinidia deliciosa) and mangosteen (Garcinia mangostana) consumed by ovariectomized rats demonstrated a beneficial effect on femoral length, weight and strength, and decreased the bone resorption marker TRAP (14). The consumption of berries has also been reported to protect from bone loss (15). Eventually, extracts, juices and fruit which are rich in polyphenols and antioxidants and have experimentally demonstrated a bone-protective action, may appear attractive alternative options for postmenopausal women who are incompliant to drug therapy.
Genome-wide studies in the management of osteoporosis
Omics studies as tools in translational research: a short historical reference
In the recent years, new methods for quantitative high-throughput analysis of genes, transcripts and proteins have developed and are applied in the area of disease genomics and in particular, osteoporosis. Researchers have established that genome wide detection of DNA low copy number changes is possible using comparative genomic hybridization (CGH) arrays. At the mRNA level, gene expression profiling is achievable through the introduction of cDNA (16) and oligonucleotide microarrays (17), which permit simultaneous analysis of thousands of genes. Real-time quantitative polymerase chain reaction (qRT-PCR) has been developed as the new standard for accurate quantification selected subsets of gene specific DNA or RNA sequences (18). Along with the genomics and transcriptomics research areas, proteomics also plays a lead role in cancer research, as a result of new and powerful analytical methods.
cDNA microarrays have provided us with a huge amount of information regarding the transcriptional status of thousands of genes. But transcription is followed by translation and post-translational modifications (PTMs) and therefore the level of mRNA does not allow someone to predict the level of protein expression. This is partly due to the PTMs and partly due to the fact that protein maturation and degradation are dynamic processes, which can dramatically alter the final amount of active protein independently of the mRNA level.
The miRNome in the mechanistics of osteoporosis
MicroRNAs (miRNAs) are small, single-stranded, non-coding RNAs, consisting of 20–25 nucleotides, whose role is post-transcriptional regulation of gene expression. miRNAs are partially complementary and bind to the 3’-untranslated region (3’-UTR) of their target messenger RNA (mRNA). They then inhibit the translation of their target mRNA or cause its degradation. Thus, miRNAs inhibit the expression of their target gene at post-transcriptional level (19). Concerning the synthesis of miRNAs, primary miRNA is transcribed in the nucleus from its gene. Afterwards, ribonuclease Drosha and protein DGCR8 process the primary miRNA to precursor miRNA. The precursor miRNA is transferred to the cytoplasm and ribonuclease Dicer processes it to mature miRNA. The passenger strand of the miRNA is ejected and degraded and the other strand—the mature miRNA—is loaded to protein Argonaute (Ago). The mature miRNA interacts with proteins Ago and GW182, binds to its target mRNA and inhibits its translation.
miRNAs are encoded by DNA sequences which are found in the genome either as separate miRNA genes or within the introns of other genes. Over 3% of human genes have been found to contain miRNA-coding sequences, while the expression of 40–90% of human protein-coding genes is regulated by miRNAs. The expression of a protein-coding gene may be regulated by more than one miRNA and each miRNA may regulate the expression of several target genes. miRNAs participate in many biological procedures, such as cell differentiation, proliferation and apoptosis, and they are involved in several diseases, including cancer, viral infections, bone metabolic diseases and autoimmune diseases (20).
In a recent study by Wang et al. (21), miR-133a was identified as a possible molecule, whose expression levels varied between patients with low BMD as compared to patients with high BMD values. Further on, microarray analysis of circulating mononuclear lymphocytes revealed that miR-133a manifested higher expression levels in patients with low BMD as compared to those with high BMD. In the same study, and in accordance to our previous comments in the previous sections, bioinformatic target gene analysis demonstrated three potential osteoclast-related target genes, CXCL11, CXCR3 and SLC39A1 of miR-133a (22). These molecules could be considered as potential biomarkers. In another recent study, miRNAs array analysis of osteoclast precursors from ten low BMD and ten high BMD patients determined upregulation of miR-422a in the low BMD patients, as compared to the high BMD patients. A more significant upregulation of miR-422a was determined in the low BMD group by qRT-PCR analysis, as compared to high BMD patients (21). From these findings, it appeared that miR-422a could be a potential biomarker underlying postmenopausal osteoporosis (21). In another recent study by Meng et al. (23), miR-194-5p was identified as a potential biomarker for postmenopausal osteoporosis. Microarray analysis from osteopenia and osteoporosis blood samples identified five miRNAs (miR-130b-3p, miR-151a-3p, miR-151b, miR-194-5p and miR-590-5p), with miR-194-5p being the only molecule with elevated expression; it was also noted to be increased in several osteoporosis-related pathways (23). Additionally, in a study by Panach et al. (24), three miRNAs (miR-122-5p, miR-125b-5p and miR-21-5p) were identified to be upregulated in blood samples of brain trauma patients compared to control samples. Some of the aforementioned results are summarized in Table 1, which includes the miRNA expression profile of osteoporotic patients based on data from the studies by Wang et al. (21), Meng et al. (23), Panach et al. (24), Cao et al. (25), Kocijan et al. (26), Seeliger et al. (27) and Kumar et al. (28).
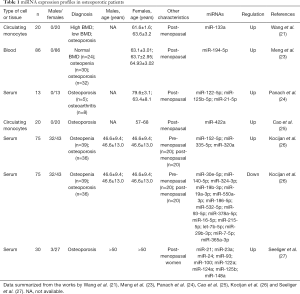
Full table
The transcriptome in the mechanistics of osteoporosis
Numerous studies have highlighted the role of gene expression and in particular of gene regulation. For example, several studies have identified some target genes of the vitamin D receptor, which included among others BsmI, ApaI, TaqI, Cdx2 and FokI. Specifically, Cdx2 polymorphism was associated with femoral neck BMD in a study of 239 osteoporotic postmenopausal women carried out by Mencej-Bedrac et al. (29). In another recent report by Gentil et al. (30), Cdx2 polymorphism did not influence BMD in osteoporotic women by itself, but actually affected the BMD response to physical activity.
Lately, osteoblast and osteoclast differentiation and maintenance have been the focal point of microarray experiments. Osteoblasts’ ability to produce and mineralize the bone matrix is well known. They differentiate from their mesenchymal precursors through an intricate procedure governed by the activation of specific transcription factors and hormones. Several transcriptional factors, such as Runx2/Cbfa1 and Osterix have been identified as key regulators of osteoblastic differentiation. Their absence has been demonstrated to result in complete lack of mineralized skeleton. Many other transcription factors are also considered important regulators of osteoblast differentiation and preservation, such as homeobox proteins, members of the AP1 family or effectors of the β-catenin/Wnt signaling pathway (31).
Patient-specific drug design for the management of osteoporosis
Based on the genomic profiling, recent advances have led to the discovery of new drug targets. In the case of osteoporosis, genome-wide studies have identified more than 100 genetic loci linked to BMD (32). In a recent study by Kemp et al. (33), 307 potentially independent single-nucleotide polymorphisms (SNPs) were reported, which could explain approximately 12% of the phenotypic variance. Interestingly, novel multiple variants were identified that showed a strong association with estimated heel ultra-sound BMD and little correlation with BMD derived by dual-energy X-ray absorptiometry (e.g., TBX1 and ZNRF3), analyzed in the genetic factors for osteoporosis study (33). The discrepancy might have originated from the difference in method and larger sample size. Furthermore, gene expression analysis of human and mouse bone cells, gene-function predictions, along with skeletal phenotyping of 120 knock-out mice, were integrated in this study. Glypican 6 (GPC6), a membrane surface proteoglycan involved in cellular growth control and differentiation, was defined here as a novel determinant of BMD (33). The GPC6 knock-out mice had shorter femurs and vertebrae than the wild-type mice, which provided evidence that GPC6 is important in bone growth and development and might serve as a target for novel treatments or prevention of fragility fractures.
The first Genome-Wide Association Study (GWAS) meta-analysis that correlated genome-wide genetic variants with lumbar spine volumetric BMD was recently undertaken (34). Lumbar spine volumetric BMD was measured in 15,275 individuals using quantitative computed tomography, and this resulted in five significant loci (i.e., WNT4, ZBTB40, TNFRSF11B, AKAP11 and TNFSF11). In addition, expression quantitative trait locus analyses of iliac crest biopsies were performed in 84 postmenopausal women, which resulted in two novel SNPs. The majority of GWASs have correlated genetic variance with the most widely used characteristic of bone health, BMD. The impact of genetic variance on the expression of genes has been analyzed in expression quantitative trait loci studies, where genetic variations are associated with the expression level of genes in hundreds of individuals. Alonso et al. (35) conducted a GWAS where genetic variance was associated with expression quantitative trait loci data from transiliac bone biopsies. A novel locus on chromosome 2q13, i.e., SNPrs10190845, was significantly associated with clinical vertebral fractures. On the basis of this bioinformatics analysis, tubulin tyrosine ligase and solute carrier family 20 member 1 (SLC20A1) were proposed as positional candidate genes of this locus. As SLC20A1 is a sodium-phosphate symporter, its role in regulating bone metabolism seems quite straightforward.
More than one tenth of the genes have a pleiotropic effect on multiple phenotypes. This means that a single gene can seemingly independently influence more than one phenotypic trait. The availability of the large-scale genomic datasets from various epidemiological studies has allowed the examination for pleiotropic effects of genetic traits on different phenotypes. The first systematic study of pleiotropy between osteoarthritis and BMD included GWAS data from GEFOS, arcOGEN and three osteoarthritis studies, which revealed 143 genetic variants with cross-phenotypes with osteoarthritis (36). In this study, SNPs in the intronic region of SMAD3 that are known to have roles in bone remodeling were confirmed to be of significant relevance for both the traits of osteoarthritis and BMD. A bivariate GWAS meta-analysis of total body lean mass and total-body-less-head BMD in 10,414 children identified genetic factors with pleiotropic effects on lean mass and BMD (37). Here, eight genetic loci correlated well with both the traits of skeletal lean mass and BMD (Table 1). Moreover, the gene SREBF1 was suggested to be an important pleiotropic.
Conclusions
Osteoporosis is a multi-factorial disease with complex and diverse pathogenesis mechanisms. Many experimental studies continue to investigate its causes, as well as its management, aiming to elucidate them, by using laboratory animals or molecular biology systems. These standardized and controlled systems offer uniform and replicable research opportunities, which may bear sounder results compared to clinical studies that apply even the strictest inclusion and exclusion criteria. Thus, “translation” of experimental research results are applied to the clinical setting. Regulation of an individual’s bone metabolism has been identified to date to require examination of many more parameters in addition to BMD, such as genes, DNA, biochemical and inflammatory markers, leading to the need for personalized therapy. The ultimate benefits of therapy—the prevention of osteoporotic fractures and maintenance of quality of life—continue to drive current research.
Acknowledgements
None.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
References
- Mahla RS. Stem Cells Applications in Regenerative Medicine and Disease Therapeutics. Int J Cell Biol 2016;2016:6940283. [Crossref] [PubMed]
- Consensus development conference. diagnosis, prophylaxis, and treatment of osteoporosis. Am J Med 1993;94:646-50. [Crossref] [PubMed]
- Kanis JA, McCloskey EV, Johansson H, et al. A reference standard for the description of osteoporosis. Bone 2008;42:467-75. [Crossref] [PubMed]
- Curtis EM, Moon RJ, Harvey NC, et al. The impact of fragility fracture and approaches to osteoporosis risk assessment worldwide. Bone 2017;104:29-38. [Crossref] [PubMed]
- Kanis JA, Johnell O, Oden A, et al. FRAX and the assessment of fracture probability in men and women from the UK. Osteoporos Int 2008;19:385-97. [Crossref] [PubMed]
- Dontas IA, Lelovas PP, Kourkoulis SK, et al. Protective effect of Sideritis euboea extract on bone mineral density and strength of ovariectomized rats. Menopause 2011;18:915-22. [Crossref] [PubMed]
- Lelovas PP, Xanthos TT, Thoma SE, et al. The laboratory rat as an animal model for osteoporosis research. Comp Med 2008;58:424-30. [PubMed]
- Jee WS, Yao W. Overview: animal models of osteopenia and osteoporosis. J Musculoskelet Neuronal Interact 2001;1:193-207. [PubMed]
- Bharti S, Rani N, Krishnamurthy B, et al. Preclinical evidence for the pharmacological actions of naringin: a review. Planta Med 2014;80:437-51. [Crossref] [PubMed]
- Sun X, Li F, Ma X, et al. The Effects of Combined Treatment with Naringin and Treadmill Exercise on Osteoporosis in Ovariectomized Rats. Sci Rep 2015;5:13009. [Crossref] [PubMed]
- Liu Y, Zuo H, Liu X, et al. The antiosteoporosis effect of icariin in ovariectomized rats: a systematic review and meta-analysis. Cell Mol Biol (Noisy-le-grand) 2017;63:124-31. [Crossref] [PubMed]
- Zhao H, Zhao N, Zheng P, et al. Prevention and Treatment of Osteoporosis Using Chinese Medicinal Plants: Special Emphasis on Mechanisms of Immune Modulation. J Immunol Res 2018;2018:6345857. [Crossref] [PubMed]
- Liang Q, Lv M, Zhang X, et al. Effect of Black Tea Extract and Thearubigins on Osteoporosis in Rats and Osteoclast Formation in vitro. Front Physiol 2018;9:1225. [Crossref] [PubMed]
- Vellapandian C, Sukumaran ES, Sivasubramanian LR, et al. A Comparative Study of Actinidia deliciosa and Garcinia mangostana in Ovariectomy-Induced Osteoporosis in Female Wistar Rats. Biomed Res Int 2017;2017:5349520. [Crossref] [PubMed]
- Hubert PA, Lee SG, Lee SK, et al. Dietary Polyphenols, Berries, and Age-Related Bone Loss: A Review Based on Human, Animal, and Cell Studies. Antioxidants (Basel) 2014;3:144-58. [Crossref] [PubMed]
- Song JY, Yu J, Chan WC. Gene expression profiling in non-Hodgkin lymphomas. Cancer Treat Res 2015;165:97-123. [Crossref] [PubMed]
- Lockhart DJ, Dong H, Byrne MC, et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nat Biotechnol 1996;14:1675-80. [Crossref] [PubMed]
- Bustin SA. Absolute quantification of mRNA using real-time reverse transcription polymerase chain reaction assays. J Mol Endocrinol 2000;25:169-93. [Crossref] [PubMed]
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004;116:281-97. [Crossref] [PubMed]
- Miao CG, Yang YY, He X, et al. New advances of microRNAs in the pathogenesis of rheumatoid arthritis, with a focus on the crosstalk between DNA methylation and the microRNA machinery. Cell Signal 2013;25:1118-25. [Crossref] [PubMed]
- Wang Y, Li L, Moore BT, et al. MiR-133a in human circulating monocytes: a potential biomarker associated with postmenopausal osteoporosis. PLoS One 2012;7:e34641. [Crossref] [PubMed]
- Meyre D, Froguel P, Horber FF, et al. Comment on: Valette et al. Melanocortin-4 receptor mutations and polymorphisms do not affect weight loss after bariatric surgery. PLOS ONE 2012; 7(11):E48221. PLoS One 2014;9:e93324. [Crossref] [PubMed]
- Meng J, Zhang D, Pan N, et al. Identification of miR-194-5p as a potential biomarker for postmenopausal osteoporosis. PeerJ 2015;3:e971. [Crossref] [PubMed]
- Panach L, Mifsut D, Tarin JJ, et al. Serum Circulating MicroRNAs as Biomarkers of Osteoporotic Fracture. Calcif Tissue Int 2015;97:495-505. [Crossref] [PubMed]
- Cao Z, Moore BT, Wang Y, et al. MiR-422a as a potential cellular microRNA biomarker for postmenopausal osteoporosis. PLoS One 2014;9:e97098. [Crossref] [PubMed]
- Kocijan R, Muschitz C, Geiger E, et al. Circulating microRNA Signatures in Patients With Idiopathic and Postmenopausal Osteoporosis and Fragility Fractures. J Clin Endocrinol Metab 2016;101:4125-34. [Crossref] [PubMed]
- Seeliger C, Karpinski K, Haug AT, et al. Five freely circulating miRNAs and bone tissue miRNAs are associated with osteoporotic fractures. J Bone Miner Res 2014;29:1718-28. [Crossref] [PubMed]
- Kumar S, Vijayan M, Bhatti JS, et al. MicroRNAs as Peripheral Biomarkers in Aging and Age-Related Diseases. Prog Mol Biol Transl Sci 2017;146:47-94. [Crossref] [PubMed]
- Mencej-Bedrac S, Prezelj J, Kocjan T, et al. The combinations of polymorphisms in vitamin D receptor, osteoprotegerin and tumour necrosis factor superfamily member 11 genes are associated with bone mineral density. J Mol Endocrinol 2009;42:239-47. [Crossref] [PubMed]
- Gentil P, Lima RM, Lins TC, et al. Physical activity, Cdx-2 genotype, and BMD. Int J Sports Med 2007;28:1065-9. [Crossref] [PubMed]
- Stains JP, Civitelli R. Genomic approaches to identifying transcriptional regulators of osteoblast differentiation. Genome Biol 2003;4:222. [Crossref] [PubMed]
- Estrada K, Styrkarsdottir U, Evangelou E, et al. Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nat Genet 2012;44:491-501. [Crossref] [PubMed]
- Kemp JP, Morris JA, Medina-Gomez C, et al. Identification of 153 new loci associated with heel bone mineral density and functional involvement of GPC6 in osteoporosis. Nat Genet 2017;49:1468-75. [Crossref] [PubMed]
- Nielson CM, Liu CT, Smith AV, et al. Novel Genetic Variants Associated With Increased Vertebral Volumetric BMD, Reduced Vertebral Fracture Risk, and Increased Expression of SLC1A3 and EPHB2. J Bone Miner Res 2016;31:2085-97. [Crossref] [PubMed]
- Alonso N, Estrada K, Albagha OME, et al. Identification of a novel locus on chromosome 2q13, which predisposes to clinical vertebral fractures independently of bone density. Ann Rheum Dis 2018;77:378-85. [Crossref] [PubMed]
- Hackinger S, Trajanoska K, Styrkarsdottir U, et al. Evaluation of shared genetic aetiology between osteoarthritis and bone mineral density identifies SMAD3 as a novel osteoarthritis risk locus. Hum Mol Genet 2017;26:3850-8. [Crossref] [PubMed]
- Medina-Gomez C, Kemp JP, Dimou NL, et al. Bivariate genome-wide association meta-analysis of pediatric musculoskeletal traits reveals pleiotropic effects at the SREBF1/TOM1L2 locus. Nat Commun 2017;8:121. [Crossref] [PubMed]
Cite this article as: Dontas IA, Lambrou GI. The impact of translational studies in the current management of osteoporosis. Ann Res Hosp 2019;3:9.


